SpatialTCR
An integrated platform for high-resolution spatial sequencing of T cell receptor repertoires
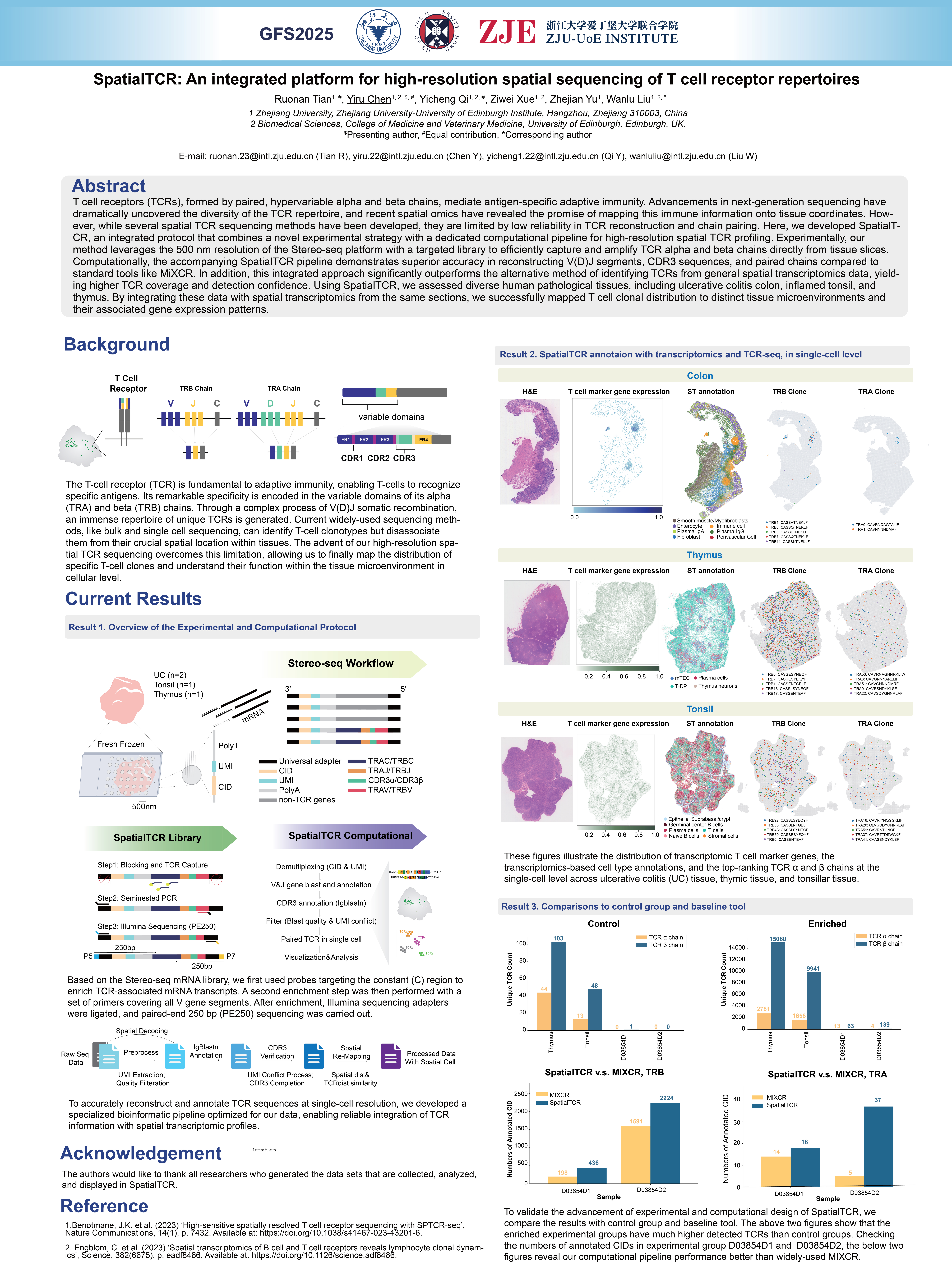
SpatialTCR: An integrated platform for high-resolution spatial sequencing of T cell receptor repertoires
Status: Independent co-first-author project; best poster award winner
Role: Co-first author, Lead computational developer
Institution: ZJU-UoE Institute, Zhejiang University
Duration: January 2025 – Present
Project Overview
SpatialTCR is an integrated experimental and computational effort for high-resolution spatial sequencing of T-cell receptor repertoires. On the computational side, the project focuses on turning raw spatial TCR data into analyzable representations that support clonotype discovery, repertoire profiling, and spatial interpretation.
My Responsibilities
As the lead computational developer, I am responsible for:
- Upstream processing: building robust workflows for raw spatial TCR sequencing data
- Algorithm development: designing computational methods for repertoire characterization in tissue context
- Analysis tooling: building downstream frameworks for TCR-omics interpretation
- Workflow integration: coordinating method development with experimental protocol design
Technical Innovation
The computational framework includes:
- sequence processing and quality control for spatial TCR reads
- statistical analysis of clonotype diversity and repertoire structure
- identification of spatially enriched TCR clonotypes
- visualization and interpretation of spatial repertoire organization
Collaborative Approach
This project is developed in close partnership with wet-lab collaborators at ZJU-UoE, with iterative feedback between protocol development and computational analysis design.
Recognition
Best Poster Presentation Award at GPB Omics & Bioinformatics Frontiers Symposium, Zhejiang, China (2025)
Presenters: Tian R†, Chen Y†#, Qi Y†, Xue Z, Yu Z, Liu W (†Co-first authors, #Presenting author)
Award-Winning Poster